All DAIR3 materials are in the GitHub Repository: https://github.com/DAIR3/DAIR3-Workshop
Unit 1: Responsible Conduct of Research (3 hours)
1.1 RCR in the Context of Biomedical Data Science
This lesson examines the sociotechnical and ethical aspects of biomedical data science. We will consider ethical issues in the responsible conduct of research that are novel to or pose new challenges in the context of biomedical data science such as reproducibility and privacy. Students will also consider biomedical data science as a sociotechnical system and define roles for themselves and other key constituents.
Learning Objectives:
- Explain novel ethical issues in responsible conduct of research for data science such as reproducibility and privacy.
- Describe the landscape of biomedical data science as a sociotechnical system and articulate roles.
Assessment Instrument:
- Compare responses to the data challenge with your peers. What issues arise?
- Why is the date and time of birth no longer recorded after 1987, only the week of birth? Discuss the privacy implications.
- Identify at least three ethical concerns when projecting underweight newborns and infant mortality by county. Consider: stigmatization of specific counties or populations; secondary use of data originally collected for other purposes; and potential biases in historical data collection.
- Who decides what data to collect, how to store it and how to access it? What biases could there be in the data (e.g., data collection in rural areas, existence of infrastructure)? What is the difference between bias and trend in these data? Provide examples.
1.2 What are Ethics? Ethical Issues in Biomedical Data Science
This lesson equips students to address ethical challenges in biomedical data science. Learners will identify strategies for ethical secondary data use, analyze engagement approaches, and develop frameworks for ethical project review, emphasizing anticipatory governance and responsible data science practices. Case studies will be used that draw on the group project selected for the 2026 cohort.
Learning Objectives:
- Differentiate between traditional bioethical, sociotechnical, and other ethical approaches to data science research and applications.
- Evaluate key ethical challenges in biomedical data science.
- Identify and formulate approaches to address ethical issues in secondary use, including anticipatory governance principles.
- Develop a framework for ethical review of biomedical data science projects.
Assessment Instrument:
- Suppose you are doing a study using NCHS vital statistics to develop an AI/ML application that will help policymakers make decisions about allocating resources for prenatal care. Who are the key stakeholders that should be involved in the design, dissemination, and/or evaluation of the application? Develop a stakeholder engagement strategy for presenting your findings to affected communities.
- You have been tasked with developing a draft governance framework for the use of predictive models in guiding decisions made by the State of Texas. Develop a framework to help the State by describing the ethical considerations and key questions and/or recommendations for approaching each ethical issue:
- Accompany your framework with a brief reflection (one paragraph) on how well it addresses key issues. Are there any issues that are missing (e.g., for individuals, communities, or populations)? How well does it anticipate future ethical challenges? What feasibility issues and resource constraints should be taken into consideration if the framework is implemented?
Unit 2: Data Management (7 hours)
2.1 Data Collection and Storage
This lesson enables learners to design robust, ethical research plans and identify the optimal data collection method for high-quality, reproducible findings.
Learning Objectives:
- Distinguish between qualitative and quantitative data.
- Apply methods for gathering, organizing, and cleaning data to ensure quality and integrity.
- Ensure ethical compliance of data collection, handling, and storage.
Assessment Instrument:
- From the Data Challenge, are the more than 100 data elements collected by the National Center for Health Statistics qualitative or quantitative?
- Were the SEER data organized and cleaned in a manner that led to your ability to reliably use them for secondary analyses? Why or why not?
- Why do you think they no longer report date and time of birth, and now only require week of birth?
2.2 Metadata – Data About Data
This lesson explores the essentials of metadata in biomedical research datasets, covering data collection methods, population, and context. Students will learn to identify high-quality metadata that supports reproducibility and distinguish it from inadequate metadata, ensuring robust and reliable research outcomes.
Learning Objectives:
- Understand standard components of metadata on biomedical science research datasets: how the data were collected, on what population, under what circumstances, etc.
- Learn to distinguish between good and bad metadata for reproducibility.
Assessment Instrument:
- List 5 critical metadata categories that researchers need to know when reproducing findings.
- From the Mathew E. Hauer article called, “Data Descriptors: Population projections for U.S. Counties by age, sex, and race,” what metadata are provided about how the population data were collected?
2.3 Data Representation
This lesson examines how data can be represented in multiple ways, highlighting that each representation impacts task efficiency. Students will learn to select optimal data representations tailored to specific research tasks, balancing ease and complexity.
Learning Objectives:
- Understand that the same data can be represented in many ways.
- Appreciate that each representation choice makes some tasks easier, but others more difficult.
- Learn how to choose a good representation for the task at hand.
Assessment Instrument:
The NCHS data are provided as a flat file with more than 100 variables. What is an alternative representation of these same data? Is the original flat file or your alternative schema more conducive to analyses, and why?
2.4.1 Data Sharing 101
This lesson introduces the principles of Open Science, focusing on the NIH Data Management & Sharing Policy’s rationale and key components. Students will explore the FAIR Guiding Principles (Findable, Accessible, Interoperable, Reusable), learning their definitions and practical examples to promote transparent and reproducible research.
Learning Objectives:
- Appreciate the foundations of Open Science
- NIH Data Management & Sharing Policy: Rationale and Key components
- FAIR Guiding Principles: Definition and examples
Assessment Instrument:
- Define rationale behind NIH Data Management & Sharing requirement.
- List 5 key components you should include in your 2-page Data Management Plan.
- List the 4 FAIR principles.
2.4.2 Data Sharing – The Reality
This lesson examines privacy and confidentiality concerns in Open Science and data sharing. Students will learn to differentiate between biomedical research types: bench science, human clinical trials, and animal models, and understand the unique data sharing implications for each, including ethical considerations and strategies to protect sensitive data while promoting transparency.
Learning Objectives:
- Learn about privacy/confidentiality concerns related to Open Science and data sharing.
- Articulate difference in types of biomedical research (bench science, human clinical trials, animal models) and what implications data sharing has for each.
Assessment Instrument:
- Create a 2-page Data Management and Sharing Plan following the NIH requirements for your analyses of the birthweight data challenge.
- Apply the FAIR principles (Findable, Accessible, Interoperable, Reusable) to your datasets.
Unit 3: Rigorous Statistical Design (5 hours)
This is a practical introduction to conducting rigorous data-driven research in a health-science setting. Our goal is that upon mastering this section, you will be able to use analytically sophisticated methods to obtain scientifically meaningful, reproducible, and innovative results in your research endeavors. We will focus here on methods for data analysis that can be utilized in the setting of population health research, leveraging official statistics or other systematically collected data with spatial and temporal structure.
This document focuses on design, strategy, and interpretation, and includes no code or numerical results. This Python notebook provides some model analyses from which you can borrow ideas as you start to implement analyses informed by the principles covered here.
3.1 Principles of Study Design for Empirical Research
In this subunit, we will consider how you can develop an understanding of the main ideas in a scientific domain in which you intend to conduct research, how to understand and communicate about the capacity of a given dataset for addressing research topics of interest, and how to develop a specific and tractable research aim.
Learning Objectives:
- You will be able to rapidly internalize the current state of scientific knowledge about a specific health-related topic. The goal here is not to master every detail that a domain specialist would know, but rather to develop fluency with the key known mechanisms, to recognize the quantitative strengths of established relationships, and to identify gaps in the current state of knowledge.
- You will be able to rapidly internalize the structure and capacity of a dataset that can be used to conduct research on a specific health-related topic. This includes identifying the units of analysis (who is being measured) and the variables or attributes (what is being measured), understanding how the units of analysis were selected, what population they represent, how the variables were measured, and what types of measurement errors may be present.
- You will be able to use appropriate terminology from epidemiology and biostatistics to discuss possible research studies relating to a given health-science domain, and that can be conducted using a provided dataset.
- You will be able to identify plausible exposures, outcomes, and control variables in a research design.
- You will be able to communicate in both speech and writing about a data-driven scientific inquiry in a health science setting. This will include strategies for effectively communicating to different audiences using precise but accessible language.
Assessment Instrument:
- In one or two sentences, state a research aim that can be addressed using the NCHS birthweight data discussed above. Your aim should be grounded in the current state of knowledge, have a limited and explicitly defined scope, and reflect a clear conceptual framework. Avoid framing the aim purely in terms of prediction accuracy or statistical methods.
- Write a roughly half-page memo to yourself providing a broader consideration of the dataset, study design, and research domain that provides the foundation for your research aim. Your memo should address: the capacity and limitations of the NCHS data for your aim; the relevant causal or mechanistic relationships (referencing the causal diagram where appropriate); the key exposures, outcomes, and potential confounders in your design; and any data quality or missingness concerns that may affect your study.
3.2 Developing an Analytic Plan
In this section, we will discuss how to develop a precise and actionable analytic plan.
Learning Objectives:
- You will be able to develop a rigorous analytic plan to address a stated scientific aim. The analytic plan should be able to be implemented using available data and should employ sophisticated and rigorous analytic methods.
- You will be able to explain the difference between conditional and marginal distributions, and identify which formal comparisons address a given research aim.
- You will be able to propose summary statistics that provide initial insight into research questions and that can capture the direction of a relationship, and the magnitude of the relationship in both relative and absolute terms.
- You will understand the different ways that an auxiliary variable can enter an analysis, such as being confounders and precision variables, and you will recognize when an auxiliary variable is unlikely to introduce confounding bias.
- You will be able to engage in a sophisticated discussion of quantitative relationships among measured quantities. This includes considering how such relationships can be assessed and how they contribute to achieving research aims.
Assessment Instrument:
- Develop a brief analytic plan (approximately one half page in length) that addresses the research aim you posed in Assessment 1, using the NCHS birthweight data. The analytic plan should be specific enough that the results would be reproducible if implemented by different researchers.
- Your plan should include: the specific regression or analytic method(s) you will employ and the rationale for selecting them; the outcome variable, key exposures, and any control or precision variables; how you will handle potential confounders, effect modification, or clustering in the data; and at least one descriptive statistic or effect size that provides initial insight into your research question, including both absolute and relative quantifications where appropriate.
- Briefly discuss any limitations of the proposed analytic approach, including assumptions that may not hold and how violations might affect your conclusions.
3.3 Bias, Causal Interpretation, and Statistical Power
Learning Objectives:
- Given a statement of research aims, a cohort study, and analysis plan for an observational study, students should be able to identify specific risks for bias, uncertainty, and non-reproducibility due to the observational nature of this study.
- In addition, students should be able to propose some elementary remedies for these challenges.
Assessment Instrument:
- Conduct a minimal power assessment that supports the analytic plan you wrote in Section 3.2. You may consider a simplified version of the analytic plan to make the power analysis more straightforward. Your assessment should include: the target parameter(s) and the effect size(s) you aim to detect; the assumed or estimated values for key quantities (e.g., residual variance, sample size, VIF); and the resulting detectable effect size or required sample size, with an interpretation of whether the available data are adequate for your aims.
- Write a short memo to yourself, no more than half a page in length, that summarizes the results of your power analysis, as well as any other reasons unrelated to power that the findings resulting from implementing your analytic plan may be misleading or spurious. Consider: potential sources of bias (e.g., confounding, informative missingness, measurement error); threats to causal interpretation given the observational nature of the data; and any limitations of the statistical methods you have chosen (e.g., model misspecification, sensitivity to distributional assumptions).
Unit 4: Unit 4: Designing Interpretable Predictive Models (5 hours)
Instructional Time: 3.5 hours
Assessment Time: 1.5 hours
This unit introduces the foundations of supervised machine learning models, with a focus on interpretability and communicating modeling decisions. Throughout the unit, the iBudget Florida Medicaid Waiver Study (Scientific Report, Volumes I and II, available as reference documents) serves as the primary worked example. The iBudget study evaluated the algorithm that Florida uses to allocate support budgets to over 35,000 individuals with developmental disabilities, building and comparing eight predictive models using real administrative data. It illustrates in a high-stakes, real-world setting every concept this unit covers: baseline model construction, interpretable feature engineering, multicollinearity diagnostics, model comparison, and stakeholder reporting. Volume II, which extends the predictive results into a resource allocation framework, is offered as supplemental reading for awareness; students are not expected to implement its methods. The parallel task throughout this unit is the 2026 Data Challenge, in which students apply the same workflow to NCHS vital statistics data to project underweight births and infant mortality by Texas county.
4.1 Foundations of Predictive Modeling
This lesson introduces supervised learning through the workflow that students will use throughout the rest of the unit: selecting a prediction target, choosing features, fitting an interpretable baseline model, and evaluating it on held-out data. The emphasis is on understanding what the model is doing, not only on obtaining a good score.
Learning Objectives:
- Distinguish supervised learning from other common data analysis tasks, and distinguish regression from classification.
- Before fitting any model, articulate the prediction problem in research terms: name the outcome variable and how it is operationalized in the dataset, identify the unit of analysis, describe the population represented, and state the intended use of the forecast.
- Build a linear regression model as an interpretable baseline and explain what its coefficients represent.
- Practice the workflow for performing a train-test split, training a model on training data, evaluating it on held-out test data, and understanding how cross-validation works in practice.
- Identify common risks in predictive modeling, including overfitting and data leakage.
Assessment Instrument:
Students will build a baseline linear regression model using the Vital Statistics dataset. They will document the outcome variable and explain how it is operationalized in the NCHS records, list the features used, report one test-set performance metric such as RMSE or MAE, and write a brief note identifying two ways data leakage or overfitting could occur in this workflow.
4.2 Interpretable Feature Engineering and Pipelines
This lesson focuses on interpretable feature engineering: creating features that better reflect patterns in the data while preserving interpretability. Students will use plots and domain knowledge to motivate simple transformations, encodings, and interactions, and will organize these steps within a small sklearn Pipeline.
Learning Objectives:
- Use visualizations to spot patterns in the data that may motivate feature engineering, such as nonlinearity, thresholds, and group differences.
- Understand when to use simple transformations, interaction terms, and common encoding strategies such as one-hot encoding.
- Build a small preprocessing and modeling Pipeline in sklearn and explain how the resulting features affect model interpretability.
Assessment Instrument:
Students will continue working with the Vital Statistics dataset and will add two or three engineered features in a small sklearn Pipeline. They will submit the pipeline structure and a short written justification for each added feature, including why it may improve model fit or interpretability.
4.3 Feature Justification, Multicollinearity, and Model Diagnostics
Building upon Section 4.2, students will learn how to decide whether a feature should remain in a model. The emphasis is on practical diagnostics and justification using correlations, multicollinearity checks, variance inflation factors, coefficient stability, and residual plots.
Learning Objectives:
- Use simple quantitative tools such as Pearson and Spearman correlation, multiple R², and variance inflation factor to identify useful, redundant, or unstable features and to recognize multicollinearity.
- Use residual plots and side-by-side model comparison to assess whether added complexity is justified.
- Explain why a feature was kept, transformed, or removed based on predictive performance, interpretability, and stability.
- Examine model performance separately for meaningful subgroups, such as geographic region or demographic category, and recognize when a model that performs well overall may perform unevenly across the population it serves.
Assessment Instrument:
Students will compare two candidate models, such as a baseline model and an expanded feature-engineered model. They will decide which model they would keep and justify the decision using at least three pieces of evidence drawn from performance metrics, multicollinearity checks, coefficient stability, or residual diagnostics. The justification should also include one observation about whether the preferred model’s performance is consistent across at least one meaningful subgroup in the data.
4.4 Model Reporting and Stakeholder Communication
This lesson focuses on communicating model performance and design decisions to external stakeholders. The goal is to turn the analytic work from Section 4.3 into a clear written summary that reports model results responsibly and in a form that aligns with the broader reporting expectations introduced earlier in the curriculum.
Learning Objectives:
- Understand how to report and compare interpretable predictive models using appropriate regression metrics such as MSE, RMSE, and MAE.
- Summarize model design choices, strengths, and limitations in language appropriate for a non-technical audience.
- Practice connecting model reporting to broader ideas of transparency and reproducibility, including the spirit of TRIPOD-style reporting.
Assessment Instrument:
Students will write a short report for a stakeholder describing their final model. The report should identify the prediction target, summarize the features included, report one or two performance metrics, and explain at least one important limitation or caveat in plain language. The report should also include one sentence connecting the model to the data challenge: what specific projection will this model contribute to the final analysis, and for which population or geography?
Unit 5: Reproducible Workflows (5.5 hours)
5.1 Goals of Reproducible Analyses
Learn the key goals and challenges of creating reproducible, transparent, and user-friendly analyses that are easy to share and reuse.
Learning Objectives:
- Awareness of key challenges and goals when creating reproducible workflows, including making analyses reproducible, user friendly, transparent, reusable, version controlled, and archived.
Assessment Instrument:
- Write in your own words a summary of the key challenges and goals when creating reproducible workflows as it applies to data generally, your own work, and the CDC dataset.
5.2 Reproducibility via Code Notebooks
Gain awareness of Markdown, Jupyter, and Quarto, and learn how these tools integrate to create clear, reproducible workflows for data analysis and reporting.
Learning Objectives:
- Awareness of Markdown, Jupyter, Quarto, and how these tools can be integrated into reproducible workflows.
Assessment Instrument:
- Create a notebook that does some simple EDA on some data. It should generate at least one plot and at least one table. Create a script to download the CDC dataset. Upload your work.
5.3 Best Practices for Reproducible Programming
Learn essential best practices for reproducible programming, including writing clear scripts and functions, avoiding magic numbers, using caching and seeding for randomness, and refactoring code to enhance clarity, reliability, and repeatability.
Learning Objectives:
- Awareness of best practices for reproducible programming including writing scripts, functions, avoiding magic numbers, caching and seeding randomness, and how to refactor code to align with these practices.
Assessment Instrument:
- Add documentation to your simple EDA notebook. Create a Makefile similar to the one shown in class (e.g., with commands to download the data, run the analysis, etc.). Upload your analysis file.
5.4 Version Control
Gain a basic understanding of Git, its advantages, and learn to perform essential tasks such as cloning repositories, committing changes, and syncing with remote repositories using push and pull commands.
Learning Objectives:
- Familiarity with Git and its benefits, and the ability to begin using it for simple tasks, including cloning, committing changes, pushing and pulling.
Assessment Instrument:
- Put the source code for a template project on GitHub. Submit the link to your GitHub page. Describe any challenges you encounter.
5.5 Containers
Gain hands-on experience with key dependency management tools (Python virtual environments, renv, and containerization), understanding their pros and cons, and develop the skills to create and run basic Docker images.
Learning Objectives:
- Familiarity with various tools for dependency management, including Python virtual environments, renv, and containerization, and their respective strengths and weaknesses. Ability to create and run simple Docker images.
Assessment Instrument:
- Put a template project into an image, and ensure that it runs as expected. Upload the Dockerfile/Containerfile and other source files to the GitHub project. Describe any challenges you encounter.
5.6 Assembling a Full Analysis Pipeline
Learn key factors in organizing an analysis pipeline and develop the skills to assemble a complete, reusable pipeline template.
Learning Objectives:
- Considerations when organizing an analysis pipeline, and the ability to assemble a full template pipeline.
Assessment Instrument:
- Create a template project. It should have a directory structure similar to the one shown in class (e.g., with a data directory, analysis directory, etc.) and include a few simple markdown files. Push the full analysis to GitHub. Describe any challenges you encounter.
Unit 6: Data Integration and Meta-Analysis (3 hours)
6.1 Key Concepts in Data Integration
This lesson provides an introduction to meta-analysis as a tool for quantitative data integration and research synthesis. Data integration arises when either raw data or summary statistics are available for many subpopulations, for the same or a similar set of outcomes and predictors. In this setting, the goal is to understand the common and distinct ways that the predictors and outcomes are distributed and are related to each other across the subpopulations. Research synthesis arises when results from a collection of research studies considering similar or identical research questions are available. The studies may differ in the populations assessed, in the study designs, or in the methods used. Combining evidence across diverse studies can yield a more accurate consensus estimate of the relationships of interest and can reveal whether there is heterogeneity among the subpopulations in how these relationships are structured.
This document focuses on design, analysis, and interpretation. It includes no code or numerical results. This Python notebook provides some model analyses from which you can borrow ideas as you start to implement analyses informed by the principles covered here.
Learning Objectives:
- Identify settings where data integration is possible and likely to be informative.
- Explain the principle of inverse variance weighting and why it is more effective than simple averaging.
- Explain the difference between uncertainty and estimation imprecision in individual studies or subpopulation-specific results, and contrast this type of variation with the idea of heterogeneity in the true values of a parameter in different studies or subpopulations.
- Explain and contrast error control via the false positive rate, the family-wise error rate, and the false discovery rate, and explain the key ideas behind the local false discovery rate approach.
- Explain the basic mathematics behind methods for pooling estimates, pooling standard errors, and combining p-values from independent sources, and identify methods for combining p-values that are robust to some forms of dependence.
Assessment Instrument:
- In one or two sentences, define a research aim that could be addressed using subpopulation estimates of birthweights, where the subpopulations are defined in a way that you specify.
- Write a roughly half-page memo to yourself identifying one or two key challenges that would arise when attempting to resolve your research aim, and indicating how these challenges could be overcome using methods discussed in this unit.
6.2 Assessing Heterogeneity
Heterogeneity in statistics is mainly used to refer to situations where the structural relationships among measured or latent quantities vary across the subpopulations being assessed. This is not variation in the observations themselves, which vary according to a probability distribution determined by measurement errors and intrinsic variation. Instead, it refers to a deeper level of variation that is governed by all of the underlying factors, often unknown, that characterize the different subpopulations.
In this section, we will discuss ways to define heterogeneity in different settings, and how to estimate it from data. The major challenge will be to distinguish the variation of the two types discussed above – the inter-individual variation driven by measurement error and individual differences, and the heterogeneity driven by structural subpopulation differences.
Learning Objectives:
- Identify opportunities to apply visualization methods from applied statistics to the settings of meta-analysis and data integration.
- Articulate the difference between statistical uncertainty and effect heterogeneity.
- Explain the basis and purpose for the intraclass correlation coefficient, and contrast it with τ², the variance of true parameter values across subpopulations.
- Explain how the law of total variation partitions total variation into the explained and unexplained fractions of variability.
- Identify at least one setting where pooling by weighted averaging is inappropriate, and explain a more appropriate way to pool evidence in this setting.
Assessment Instrument:
- In one or two sentences, propose a brief research aim for the NCHS data centering on heterogeneity.
- Write a brief reflection discussing the approach you would employ to resolve your research aim, and discussing at least one potential challenge that is likely to arise and how you would overcome it.
6.3 Modern and Robust Approaches to Evidence Combination
This section discusses more modern and advanced methods for data integration. These approaches are more conceptually advanced than the methods discussed earlier, but may yield deeper results, especially in more challenging settings.
Learning Objectives:
- Articulate the purpose and main idea behind the jackknife empirical likelihood approach in a data integration setting.
- Explain and contrast the definitions of p-values and E-values, and identify some advantages and effective uses for each.
- Explain the rationale for partitioning variance using estimated variance components in a meta-analysis setting, and explain the meanings of explained variation, random main effects, and idiosyncratic variation.
Assessment Instrument:
- Choose one of the three approaches discussed in this unit (jackknife empirical likelihood, E-values, or variance component analysis), and write a persuasive paragraph addressed to a clinical researcher, aiming to explain in a non-technical way the basis of the approach, and proposing a possible way for them to take advantage of this newer and more advanced methodology in their research.
Unit 7: Large Language Models in Biomedical Research (5 hours)
This unit prepares participants to use large language model (LLM) based AI agents as rigorous, accountable tools in biomedical data science research. The unit is load-bearing: its deliverables are components of the 2026 Data Challenge final report. Students work with the same NCHS vital statistics database used throughout the bootcamp, using an LLM ensemble to assist with data navigation, methodological decision-making, grant review, and manuscript preparation, while maintaining human accountability for every output.
Three principles organize the unit. First, invalidation (a framework broader than “hallucination” that encompasses factual, logical, normative, and structural breaches) is mathematically inevitable in any LLM-generated output; human verification is therefore not optional. Second, cross-agent critique reduces invalidation rates below what any single model achieves, but introduces its own ethical tradeoffs that must be actively managed. Third, any sociodemographic variable whose encoding has changed across the historical record carries risks of structural and normative invalidation when queried without contextual grounding.
7.1 The Ethics of AI Agents
This lesson examines the ethical dimensions of using LLM-based AI agents in biomedical data science. Unlike single-turn chatbot interactions, agentic workflows involve planning, tool use, persistent memory, and multi-agent coordination, substantially expanding the ethical surface area of AI-assisted research. Students develop a framework for identifying and classifying invalidation risks, analyze the specific ethical challenges posed by historical vital statistics data with inconsistently encoded racial and ethnic categories, and produce a Research AI Governance Protocol that governs their own use of LLM tools throughout Sections 7.2 and 7.3.
The lesson proceeds in four instructional blocks:
- Agent architecture and expanded ethical surface (15 min). What distinguishes an AI agent from a chatbot: planning, tool use, memory, and multi-agent coordination. Each capability adds a distinct ethical surface. The discursive network framework explains why agent-generated statements that enter manuscripts acquire authority independent of their accuracy, making verification a structural responsibility rather than an optional quality check.
- Invalidation: a taxonomy broader than hallucination (15 min). The four types of invalidation (factual, logical, normative, and structural) are introduced using NCHS-specific examples. Factual: an LLM names a variable not present in the DVS schema as a named field. Logical: an LLM recommends a rate denominator inconsistent with the research question’s population definition. Normative: an LLM interprets racial disparities in birthweight as reflecting biological differences between racial groups rather than the consequences of racism. Structural: an LLM queries APGAR score columns for records before 1977, when those columns did not exist in the flat file. Each type requires a distinct verification strategy.
- Sociodemographic consciousness in biomedical data science (15 min). We explore how many sociodemographic variables are social construct with no valid biological basis, but their reification in law, medicine, and data systems makes it real in its consequences. Birth outcomes differ by recorded race not because of intrinsic racial biology but because of racism: differential access to prenatal care, chronic stress from discrimination, environmental exposures in historically redlined neighborhoods, and differential clinical treatment.
The specific NCHS data students are working with encodes race through a system that changed across the study period. From 1969 through 1977, race was assigned from a fixed set (White, Black, American Indian, Chinese, Japanese, Other), and for parents of different recorded races the child’s race followed the mother’s. Assignment rules were revised in 1978, inconsistently across states. The “Hispanic” ethnic category appeared late in the period and was collapsed into “White” in some years. The child_race_recode_3 field changed its encoding midway through the study period. Any LLM query about race in this dataset risks producing normative invalidation by treating these categories as stable, comparable, and biologically grounded across the full 1969–1986 span. - Ethics checklist and governance protocol construction (20 min). The seven risk categories from the ethics framework (epistemic, accountability and authorship, provenance and reproducibility, confidentiality, bias and fairness, security, and sustainability) are reviewed with emphasis on the tradeoffs specific to this bootcamp context. Multi-agent critique tradeoffs: privacy amplification, epistemic homogenization, compute burden, and attribution diffusion. Students construct the Research AI Governance Protocol that governs their subsequent work.
Learning Objectives:
- Define an AI agent in terms of planning, tool use, memory, and multi-agent interaction, and explain how each capability expands the ethical surface area relative to single-turn LLM use.
- Apply the four-type invalidation taxonomy (factual, logical, normative, and structural) to concrete scenarios involving the NCHS vital statistics database, and identify the appropriate verification strategy for each type.
- Articulate why sociodemographic variables historical US vital statistics data often reflect social constructs whose reification has real health consequences, and explain the specific coding inconsistencies in the NCHS 1969–1986 dataset that make naive racial comparisons normatively invalid.
- Construct a Research AI Governance Protocol specifying provenance logging, invalidation classification, sociodemographic variable handling, escalation thresholds, and a compliant disclosure statement for the team’s subsequent AI-assisted work.
Assessment Instrument:
Item 1 — Invalidation Taxonomy Application (in-session, 8 minutes): The instructor presents four NCHS-specific LLM output scenarios, one per invalidation type. For each scenario, teams identify: (a) the invalidation type, (b) why it constitutes that type and not another, and (c) the verification method that would detect and correct it. Scenarios are drawn from realistic ensemble outputs on the vital statistics database.
Item 2 — Research AI Governance Protocol (in-session construction, 10 minutes; referenced throughout Sections 7.2 and 7.3): Each team produces a one-page governance protocol containing all five required components:
- Provenance logging convention: how prompts, model identifiers, and timestamps will be recorded for every ensemble interaction.
- Invalidation log format: the template for recording invalidation type, description, and resolution for each detected LLM failure.
- Sociodemographic variable handling clause: specific commitments about how sociodemographic variables with historically unstable encoding will be treated in LLM queries. LLM queries involving sociodemographic variables must inject the relevant coding history as context; temporal comparisons of category rates require explicit justification; normative claims about disparities must be flagged for human review.
- Escalation threshold: the conditions under which LLM output will not be used without independent verification or correction (e.g., any factual claim about NCHS variable definitions; any normative statement about population health disparities; any generated code before execution).
- Draft disclosure statement: a single sentence in the form specified by the ethics framework, naming tools and tasks, to be finalized at the end of Section 7.3.
The governance protocol is a live document. It is updated throughout Sections 7.2 and 7.3 as invalidations are encountered and resolved, and its disclosure statement is finalized as the unit’s final deliverable.
7.2 AI Agents for Technical Tasks
This lesson teaches students to use an LLM ensemble as a rigorous technical collaborator on analytical tasks grounded in the NCHS vital statistics database. The session has four sequential blocks. Block A derives the mathematical basis for multi-agent consensus and produces the agent-count commitments that govern subsequent activities. Blocks B, C, and D form an analytical chain: outputs from each block feed the next, and their collective products are load-bearing components of the final data challenge report.
Students enter this session with a completed model report from Unit 4, which documents a predictive model for the data challenge outcome (underweight birth rate or infant mortality rate by Texas county) including the feature selection memo, model comparison table, residual diagnostics, and limitations section. This report is the primary document the LLM ensemble will be asked to interrogate. The governance protocol from Section 7.1 is active from the start of this session; every ensemble interaction concerning the Unit 4 model is logged in the invalidation log.
The governing framework is the Flaws-of-Others (FOO) algorithm, in which multiple agents independently produce outputs, each critiques the others’ responses, and a harmonizer synthesizes the critiques into a refined result. The minimum number of mutually detecting agents required to reduce the long-run false-statement share below a tolerance ε is:
nmin
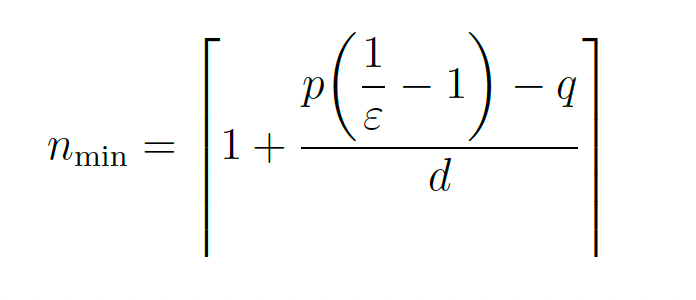
where p is the per-agent invalidation rate, q is the internal self-repair rate (q = 0.05 reference value), and d is the cross-detection probability (d = 0.19 reference value). Students use this formula to make defensible, quantitative decisions about ensemble size for each activity.
Block A — FOO Framework: Derivation and Calibration (20 minutes instructional)
The derivation proceeds from first principles. A single agent has an effective invalidation rate p (combining intrinsic corruption and fabrication). Internal self-repair operates at rate q. Each additional cross-checking agent adds detection hazard d to the effective correction rate. The steady-state false-statement share with n mutually detecting agents is:
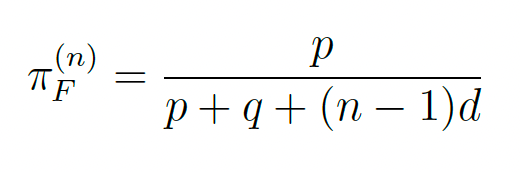
Setting πF(n) ≤ ε and solving for n yields Eq. (7.1). The invalidation floor (proved formally in the underlying theory) establishes that some error rate is mathematically inevitable in any finite-loss LLM; the FOO architecture is the engineering response to this impossibility result.
Students compute nmin for three task categories they will encounter in this session using the reference values above:
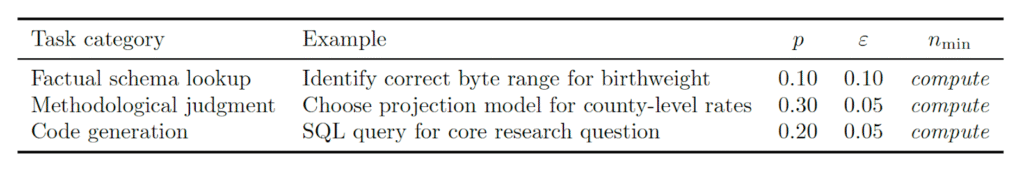

The computed values are entered into the governance protocol as the team’s agent-count commitments for Blocks B, C, and D.
Block B — The Texas Births Problem (35 minutes)
Before the bootcamp, participants were asked to count live births in Texas in 1969 to mothers residing in Texas. Teams compare their answers. Historically, different correct implementations of this query return different counts. The task is to diagnose the source of disagreement using the LLM ensemble.
Teams query the ensemble with the NCHS data dictionary and ask it to: (a) identify every field relevant to the filter, (b) flag any ambiguities in the dictionary language, and (c) write a query implementing the most defensible interpretation. Teams execute the generated query and compare counts across the class.
The deliverable is annotated, executable code (Python or SQL) that implements the correct filter with comments explaining each disambiguation decision. This code is a candidate component of the reproducible pipeline developed in Unit 5.
Model report critique (15 min). Each team submits their Unit 4 model report to the ensemble and prompts it to identify any claim in the methods section that is (a) not reproducible from the written description alone, (b) in tension with a stated limitation, or (c) likely to require verification against the NCHS data dictionary. Teams record each flagged claim in the invalidation log with an invalidation type classification from Section 7.1. For each factual or structural invalidation identified, teams verify the ensemble’s critique against the Unit 4 model code before accepting or rejecting it. Teams that find no invalidations in their Unit 4 report should examine whether the ensemble is producing false-negative outputs (a structural invalidation of the ensemble itself) and document their reasoning.
Block C — Sociodemographic Variable Integrity: Instructor-Led Exploration (15 minutes)
Building directly on Section 7.1, the instructor demonstrates how LLM behavior differs across three sociodemographic variables whose encoding changed across the NCHS study period. For each variable the instructor submits a common query to the ensemble, first without contextual grounding and then with the relevant coding history injected, and the class observes the difference in output.
The three demonstrations pursue the same comparative question: does the ensemble recognize that the variable is historically unstable, and does it flag its own normative or structural claims without prompting? Race is the variable most likely to surface normative invalidation; maternal education is most likely to surface structural invalidation arising from changes in coding categories; geographic identifiers are most likely to surface factual invalidation arising from county boundary or code revisions. Together they illustrate that the retrieval-augmented mitigation strategy from Section 7.1 generalizes to any variable with a documented encoding history.
Students take notes and update the sociodemographic variable handling clause in their governance protocol based on what they observe. The take-home deliverable (Assessment Item 5) asks them to apply the same comparative analysis independently to their own data challenge variables.
Block D — Methodological Decision Support (60 minutes)
Teams use the ensemble to interrogate the model family choice documented in their Unit 4 report. The core prompt pattern is: present the ensemble with the Unit 4 feature set, the outcome distribution description, and the model comparison table, then ask it to (a) identify which alternative model from the comparison was most appropriate given the stated distributional properties of the outcome, and (b) identify any model family that was not evaluated in the Unit 4 comparison but would be worth considering given the data structure. Teams apply Eq. (7.1) to determine the appropriate ensemble size for this task. Every ensemble recommendation is logged and verified against the Unit 4 model diagnostics before being accepted. If the ensemble recommends a model family that the student already evaluated and rejected in Unit 4, teams document whether the ensemble’s reasoning accounts for the diagnostic evidence that motivated the rejection.
Learning Objectives:
- Derive the FOO minimum-agent formula from the steady-state false-share equation and compute nmin for tasks of varying epistemic difficulty, applying the result to make quantitative ensemble-size decisions.
- Use an LLM ensemble to navigate an ambiguous biomedical data dictionary, verify outputs against the authoritative database schema, and produce annotated executable code that implements a defensible query with all disambiguations documented.
- Recognize temporal instability and normative invalidation risk across sociodemographic variables in the NCHS dataset, using sociodemographic variables and apply retrieval-augmented mitigation strategies consistently across all three.
- Produce a structured methodological consensus memo evaluating competing projection approaches, with FOO-calibrated confidence levels, a synthesis recommendation, and a retrospective comparison to the analytic plan developed in Unit 3.
Assessment Instrument:
Item 3 — Verified Data Query: Texas Birth Count (Block B deliverable): Annotated, executable code implementing the correct Texas birth count filter. The annotation must explain: (a) each filter field used and why, (b) each source of ambiguity in the data dictionary and how it was resolved, (c) the count produced, and (d) at least one invalidation from the ensemble interaction, logged in the governance protocol format.
Item 4 — Invalidation Log (cumulative across Blocks B, C, and D, and updated through Section 7.3): A structured log of all LLM invalidations encountered during Unit 7. Each entry must record: invalidation type (factual, logical, normative, or structural), a one-sentence description of the failure, the resolution applied, and the section in which it occurred. The log must contain at least six entries across the full unit (minimum two from Section 7.2). If a particular invalidation type does not appear in a team’s experience, this must be explicitly documented with a note on what would have caused it to appear.
Item 5 — Sociodemographic Variable Integrity Memo (take-home, due before Session 7.3): A written memo of three to four paragraphs treating sociodemographic variables comparatively. For each variable the memo must specify: (a) which coding or classification changes affect the variable across the study period and their source in the data dictionary; (b) which temporal comparisons are analytically valid and which are not; (c) how the team’s LLM queries will be structured to mitigate normative or structural invalidation specific to that variable; and (d) the updated sociodemographic variable handling clause for the governance protocol, incorporating all variables.
Item 6 — Methodological Consensus Memo (Block D deliverable): A structured memo containing: (a) a comparison table with one row per projection approach, recording ensemble consensus level (High/Medium/Low), key assumptions, major risks, and divergence points; (b) the team’s synthesis recommendation with justification; (c) the FOO analysis, which approach generated the most divergent ensemble responses and what epistemic difficulty that signals; and (d) a retrospective comparison to the analytic plan designed in Section 3.1, noting whether the consensus confirms, complicates, or challenges the earlier design choices.
7.3 AI Agents for Grant Review and Manuscript Preparation
This session transfers the technical skills developed in Section 7.2 to the most career-relevant applications of LLM-assisted research: grant review and manuscript preparation. Students work with canonical NIH grant proposals that include their actual panel summary statements, providing ground-truth verification for ensemble outputs. Students also prepare a Specific Aims page for a research proposal using Jackson Heart Study (JHS) data, applying the framework from Section 7.1 to the framing of health disparity research.
The governance protocol from Section 7.1 remains active. The special confidentiality concern of Section 7.3A is emphasized: federal agencies prohibit uploading unpublished proposal text to unapproved AI systems for grant review purposes. The proposals used here are canonical public examples; students are explicitly warned that this constraint applies to any future peer review panel work. The invalidation log from Section 7.2 is updated throughout.
Block A — NIH Grant Review with Ground-Truth Comparison (60 minutes)
Each team selects one canonical NIH grant proposal from the provided collection. These proposals are publicly available examples accompanied by their actual summary statements from review panels.
Prompt design (10 min). Teams design separate prompts for each of the five NIH review criteria: Significance, Investigator(s), Innovation, Approach, and Environment. Each criterion requires different domain reasoning; a single prompt for all five produces superficial output. Teams apply Eq. (7.1) per criterion, recognizing that Approach typically warrants a higher agent count than Environment.
Ensemble review (25 min). Teams run the consensus pipeline across all five criteria. The governance protocol is active; every ensemble interaction is logged.
Comparison analysis (20 min). Teams compare the ensemble’s consensus review to the actual panel summary statement across all five criteria, recording: consensus score, panel score, strengths identified by each, divergence type (missing context, conservative bias, detected concern missed by panel, domain-norm gap), and agreement level.
Critical synthesis (5 min). Teams identify three empirical categories from their comparison data: what the ensemble reliably detected, what it missed systematically, and what varied case by case. This analysis informs a critical reflection on appropriate LLM use in grant preparation.
Block B — JHS Specific Aims: Ensemble-Assisted Drafting (60 minutes)
The coordinator for JHS activities provides: a specific JHS research question, the relevant JHS variables, and two to three published JHS papers as grounding context.
Prompt construction (10 min). Teams design a structured prompt that injects JHS context as a system prompt. The task framing is explicit: not “write my Specific Aims” but rather “given this JHS research question, this prior work, and these three candidate aims, evaluate which aim structure is most scientifically defensible and propose the structure with highest consensus confidence, flagging any claims about JHS data or methods that require independent verification against the provided literature.”
Ensemble drafting (25 min). Teams run the FOO pipeline with JHS context. Every LLM claim about JHS data, published results, or methods is flagged for verification against the provided papers and logged in the invalidation log.
Human revision and sociodemographic framing review (20 min). Teams revise the LLM draft, correcting all logged invalidations. A mandatory step: teams explicitly evaluate whether the drafted Specific Aims treats sociodemographic factors as social determinants of health operating through identifiable structural pathways rather than as fixed biological or demographic properties of the study population. The review must confirm that the framing aligns with current NIH policy on sociodemographic variables in grant applications. Each revision is documented in the invalidation log.
Disclosure statement finalization (5 min). The draft disclosure statement from Section 7.1 is revised to reflect the actual scope and nature of AI assistance across all three sections. Teams select the appropriate disclosure level (minimal, standard, or detailed) and produce a finalized statement suitable for inclusion in a peer-reviewed publication.
Learning Objectives:
- Design criterion-specific prompts for LLM-assisted NIH grant evaluation; compare ensemble outputs to actual expert panel judgments; and characterize empirical patterns of agreement, divergence, and systematic miss.
- Construct an ensemble-assisted Specific Aims draft for a JHS research proposal, verifying all LLM claims against published JHS literature and ensuring that sociodemographic factors are framed as social determinants of health operating through structural pathways rather than as fixed biological or demographic properties.
- Produce a finalized disclosure statement appropriate for inclusion in a peer-reviewed publication, accurately reflecting the scope of AI assistance and maintaining full human accountability for all outputs.
7.3.2 Assessment Instrument:
Item 7 — NIH Grant Review Analysis (Block A deliverable): A structured comparison document containing: (a) the comparison table across all five NIH criteria with scores, identified strengths, divergence types, and agreement levels; (b) three empirical categories (reliable detections, systematic misses, and variable outcomes) derived from the comparison data with at least one concrete example per category; and (c) a one-paragraph critical reflection on appropriate LLM use in grant preparation, grounded in the evidence from the comparison rather than prior assumptions.
Item 8 — JHS Specific Aims Draft (Block B deliverable): A draft Specific Aims page produced through the ensemble pipeline and revised by the team, with: (a) all LLM invalidations documented in the invalidation log with verification sources cited; (b) a written framing review note confirming that sociodemographic factors are framed as social determinants of health operating through structural pathways are treated consistently with the framework from Section 7.1, and that the overall framing is consistent with NIH policy; and (c) a brief annotation for each aim identifying which elements are human-authored and which were substantially shaped by ensemble output, consistent with the governance protocol.
Item 9 — Final Disclosure Statement (Block B deliverable, unit-closing): A single, finalized disclosure statement appropriate for inclusion in a methods section or acknowledgments section of a peer-reviewed publication. The statement must: name the tools and approximate versions used; specify the tasks for which AI assistance was employed across the full unit; describe the verification procedures applied; and include an explicit accountability statement affirming that all claims, analyses, and conclusions were reviewed and affirmed by the team, who take full responsibility for the work. The statement must also specify whether and how the LLM ensemble was used to review or revise the predictive model report produced in Unit 4, including a description of the verification procedure applied to ensemble critiques of that report. The statement must be consistent with the completed invalidation log.
Unit 8: Jackson Heart Study (3 hours)
8.1 Introduction to the Jackson Heart Study
This lesson provides an overview of the Jackson Heart Study (JHS), focusing on its design, data collection, and variable interpretation. Students will describe the JHS’s purpose, population, and exam structure, summarize clinical, survey, and genetic data collection methods across study phases, and learn to use JHS codebooks to identify variables for research questions.
Learning Objectives:
- Describe the JHS Study Design: Explain the purpose, population, and structure of the JHS, including its major exams.
- Summarize Data Collection Methods: Identify the types of data collected (e.g., clinical, survey, genetic) and the methods used in different study phases.
- Interpret Key Variables and Codebooks: Understand how to use JHS codebooks to find variables relevant to specific research questions.
Assessment Instrument:
- Describe in your own words the purpose of the JHS, cohort characteristics, and exam waves.
- Describe how JHS collects specific data (e.g., CAC scores, lipid tests, etc.) and potential biases in data collection.
- Answer the following question: “Association between hysterectomy and cardiovascular disease in the JHS, adjusting for covariates” by locating relevant variables in the JHS codebook. Describe the variables chosen and their rationale.
8.2 The Process of Manuscript Development in the Jackson Heart Study
This lesson covers the process for requesting and obtaining JHS data. Students will learn to navigate data access procedures, including submitting manuscript or ancillary study proposals, completing data use agreements, and addressing ethical considerations to ensure responsible use of JHS data.
Learning Objectives:
- Explain Data Access Procedures: Describe the process for requesting and obtaining JHS data, including data use agreements and ethical considerations.
Assessment Instrument:
- In your own words, enumerate the steps involved in the process for developing a JHS manuscript.
- Using the information acquired from the lecture, draft a mock JHS manuscript proposal, using the Manuscript Proposal Form provided and the sample manuscript proposal.
